For the Regularized Discriminant Analysis Cross Validation, we need to compute SVD for each pair of ((lambda, gamma)), and the factorization result will be feed to the downdating algorithm to obtain leave one out variance-covariance matrix (hat{Sigma}_{kv}(lambda, gamma)). The code is a direct modification of my original SVD macro @here. It is much more than simply put “BY-” statement in PROC PRINCOMP, but still easy to do. I used the within class CSSCP from the Iris data shipped with SAS and cross verified with R. %CallR macro is used here.
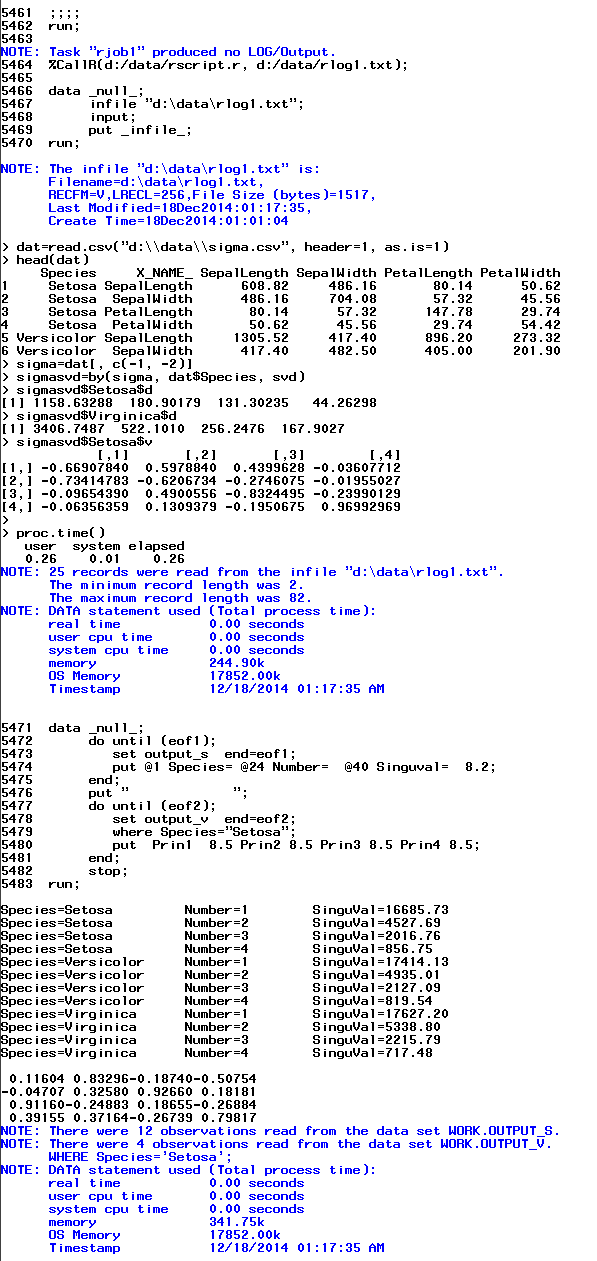
The screenshot below shows the SAS macro generates the same result as from R. Here is the test code.
dm log 'clear';
proc discrim data=sashelp.iris noclassify noprint
outstat=iris_stat(WHERE=(_TYPE_ = "CSSCP" & Species ^= " "))
;
class Species;
var SepalLength SepalWidth PetalLength PetalWidth;
run;
%svd_bystmt(sigma, output_v, output_s, output_u, &covars, Species, , nfac=0);
data _null_;
file "d:datasigma.csv" dlm=",";
set iris_stat;
if _n_=1 then do;
put 'Species' ','
'_NAME_' ','
'SepalLength' ','
'SepalWidth' ','
'PetalLength' ','
'PetalWidth';
end;
put Species _NAME_ SepalLength SepalWidth PetalLength PetalWidth;
run;
%RScript(d:datarscript.r)
cards4;
dat=read.csv("d:\data\sigma.csv", header=1, as.is=1)
head(dat)
sigma=dat[, c(-1, -2)]
sigmasvd=by(sigma, dat$Species, svd)
sigmasvd$Setosa$d
sigmasvd$Virginica$d
sigmasvd$Setosa$v
;;;;
run;
%CallR(d:/data/rscript.r, d:/data/rlog1.txt);
data _null_;
infile "d:datarlog1.txt";
input;
put _infile_;
run;
data _null_;
do until (eof1);
set output_s end=eof1;
put @1 Species= @24 Number= @40 Singuval= 8.2;
end;
put " ";
do until (eof2);
set output_v end=eof2;
where Species="Setosa";
put Prin1 8.5 Prin2 8.5 Prin3 8.5 Prin4 8.5;
end;
stop;
run;
%macro SVD_bystmt(
input_dsn,
output_V,
output_S,
output_U,
input_vars,
by_var,
ID_var,
nfac=0
);
%local blank para EV USCORE n pos dsid nobs nstmt
shownote showsource nbylevel bystmt;
%let shownote=%sysfunc(getoption(NOTES));
%let showsource=%sysfunc(getoption(SOURCE));
options nonotes nosource;
%let blank=%str( );
%let EV=EIGENVAL;
%let USCORE=USCORE;
%if &by_var eq &blank %then %do;
%let bystmt = ␣
%let nbylevel = 1;
%end;
%else %do;
%let bystmt = by &by_var;
%end;
%let n=%sysfunc(countW(&input_vars));
%let dsid=%sysfunc(open(&input_dsn));
%let nobs=%sysfunc(attrn(&dsid, NOBS));
%let dsid=%sysfunc(close(&dsid));
%if &nfac eq 0 %then %do;
%let nstmt=␣ %let nfac=&n;
%end;
%else %do;
%let x=%sysfunc(notdigit(&nfac, 1));
%if &x eq 0 %then %do;
%let nfac=%sysfunc(min(&nfac, &n));
%let nstmt=%str(n=&nfac);
%end;
%else %do;
%put ERROR: Only accept non-negative integer.;
%goto exit;
%end;
%end;
/* calculate U=XV/S */
%if &output_U ne %str() %then %do;
%let outstmt= out=&output_U.(keep=&by_var &ID_var Prin:);
%end;
%else %do;
%let outstmt=␣
%end;
/* build index for input data set */
proc datasets library=work nolist;
modify &input_dsn;
index create &by_var;
run;quit;
%let options=noint cov noprint &nstmt;
proc princomp data=&input_dsn
/* out=&input_dsn._score */
&outstmt
outstat=&input_dsn._stat(where=(_type_ in ("&USCORE", "&EV"))) &options;
&bystmt;
var &input_vars;
run;
/*
need to check if the by_var is Char or Numeric, then set the
format accordingly
*/
data _null_;
set &input_dsn;
type=vtype(&by_var);
if type="C" then
call symput("bylevelfmtvar", "$bylevelfmt");
else
call symput("bylevelfmtvar", "bylevelfmt");
run;
data __bylevelfmt;
set &input_dsn._stat end=eof;
&bystmt;
retain _nbylevel 0;
retain fmtname "&bylevelfmtvar";
if first.&by_var then do;
_nbylevel+1;
start=&by_var;
label=_nbylevel;
output;
keep label start fmtname ;
end;
if eof then call symput("_nbylevel", _nbylevel);
run;
proc format cntlin=__bylevelfmt;
run;
%put &_nbylevel;
%if &output_V ne &blank %then %do;
proc transpose data=&input_dsn._stat(where=(_TYPE_="&USCORE"))
out=&output_V.(rename=(_NAME_=variable))
name=_NAME_;
&bystmt;
var &input_vars;
id _NAME_;
format &input_vars 8.6;
run;
%end;
/* recompute Proportion */
%if &output_S ne %str() %then %do;
data &output_S;
set &input_dsn._stat ;
&bystmt;
where _TYPE_="EIGENVAL";
array _s{*} &input_vars;
array _x{&nfac, 3} _temporary_;
Total=sum(of &input_vars, 0);
_t=0;
do _i=1 to &nfac;
_x[_i, 1]=_s[_i];
_x[_i, 2]=_s[_i]/Total;
if _i=1 then
_x[_i, 3]=_x[_i, 2];
else
_x[_i, 3]=_x[_i-1, 3]+_x[_i, 2];
_t+sqrt(_x[_i, 2]);
end;
do _i=1 to &nfac;
Number=_i;
EigenValue=_x[_i, 1]; Proportion=_x[_i, 2]; Cumulative=_x[_i, 3];
S=sqrt(_x[_i, 2])/_t; SinguVal=sqrt(_x[_i, 1] * &nobs/&_nbylevel);
keep &by_var Number EigenValue Proportion Cumulative S SinguVal;
output;
end;
run;
%end;
%if &output_U ne %str() %then %do;
data &output_U;
array _S{&_nbylevel, &nfac} _temporary_;
if _n_=1 then do;
do until (eof);
set &output_S(keep=&by_var Number SinguVal) end=eof;
where Number<=&nfac;
%if &by_var eq &blank %then %do;
_row=1;
%end;
%else %do;
_row = input(put(&by_var, &bylevelfmtvar..), best.);
%end;
_S[_row, number]=SinguVal;
if abs(_S[_row, number]) < CONSTANT('MACEPS') then _S[_row, j]=CONSTANT('BIG');
end;
end;
set &output_U;
array _A{*} Prin1-Prin&nfac;
%if &by_var eq &blank %then %do;
_row=1;
%end;
%else %do;
_row = input(put(&by_var, &bylevelfmtvar..), best.);
%end;
do _j=1 to dim(_A);
_A[_j]=_A[_j]/_S[_row, _j];
end;
keep &by_var &ID_var Prin1-Prin&nfac ;
run;
%end;
proc datasets library=work nolist;
modify &input_dsn;
index delete &by_var;
run;quit;
%exit:
options &shownote &showsource;
%mend;

Read more →
